Scientists Sleuth Out the Mysteries of Dynamic Protein Droplets Formed Within Cells
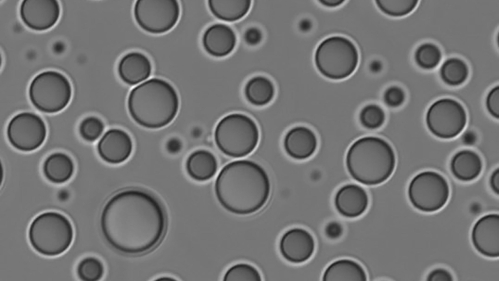

By studying novel biomolecular structures, Rutgers–New Brunswick scientists hope to better understand their role in health and disease
Benjamin Schuster enjoys solving biomedical mysteries.
An assistant professor in the Department of Chemical and Biochemical Engineering in the Rutgers University–New Brunswick School of Engineering, Schuster is enmeshed in exploring a host of frontier fields – from synthetic biology, which involves redesigning organisms for useful purposes, to “soft matter,” squishy materials from paint to polymers that are easily affected by external forces.
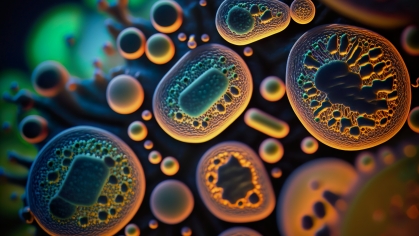
Schuster presently is knee-deep in efforts to understand the rules governing the creation of mysterious liquid protein droplets found deep within cells, alternatively known as biomolecular condensates or membrane-less organelles.
In a Nature Chemistry paper he wrote with his doctoral student, Mayur Barai, and colleagues from Texas A&M University and the University of Delaware, Schuster described experiments revealing new insights into the behavior of proteins that produce the droplets.

Schuster discussed the findings, the importance of the droplets and what this field of research may mean for the future treatment of neurodegenerative diseases.
What, exactly, are these mysterious, tiny liquid droplets – you say they are called either biomolecular condensates or membrane-less organelles?
There’s a whole world of activity going on in each living cell. The hubs of activity in our cells are known as organelles, which perform vital functions.
Most people have heard of some of the more important organelles, like the nucleus, which stores genetic information, and the mitochondria, which produce chemical energy.
Curiously, some organelles defy traditional convention. Instead of being enclosed within a protective membrane, they are without surrounding membranes and exist in the form of distinct liquid droplets. The rules that govern the formation of these droplets, a process called “liquid-liquid phase separation,” is a new and hotly pursued area of research.
The droplets were discovered in 2009 in the cells of a roundworm, C. elegans. They were strangely dynamic. They could hold their integrity, much like oil droplets in water. They could drip and join to form bigger granules. Or they could dissolve away and then later condense again, like dewdrops. This discovery changed scientists’ thinking on how cells are organized.
Why are these special droplets so intriguing?
The study of these droplets is a central part of an exciting, and fairly new, field of research that has attracted interest from biologists, chemists, physicists and engineers. Cells are very complex and crowded with molecules, and compartments such as those exhibited by the droplets help cells stay organized.
This research is important because studies are showing that these droplets play important roles inside cells, including how cells respond to stress. It's possible that they can contribute to the development of neurodegenerative diseases such as Alzheimer’s and Parkinson’s when they are converted from a liquid to a solid. It’s also been hypothesized they play a role in cancer and in replication of viral infections, including COVID.
We are very interested in how these discoveries can eventually be turned into advances in biomedicine and biotechnology. It all starts with understanding the code of the proteins and other molecules that make up these cellular droplets.
Can you explain why understanding more about the proteins that create the droplets will lead you to insights about the droplets themselves?
The droplets are formed by a class of proteins known as intrinsically disordered proteins, which lack a fixed three-dimensional structure. In attempting to understand their nature, we’re trying to decipher a foreign language.
Imagine that you only know Japanese, and no English, and a friend gives you a copy of a Harry Potter novel. After spending some time with the book, you might notice that some letters are seemingly more important, for instance the letter “o” is more common than the letter “z.” If you study the pages some more, you might notice some letters that tend to interact with each other, for instance “q” is almost always followed by “u.” From there, it’s still a long way until you can fluently read and understand the book, but you’ve at least made a good start.
In our case, we’re not studying spoken language, but rather trying to decipher the language of proteins. Proteins are made up of a string of amino acids, which you can think of as letters in a sentence, and we’re trying to understand which letters are most important and how they interact with each other. The unique combination of letters and their interactions in a protein sequence gives the protein its properties and functions.
In particular, we’re focusing on a subset of intrinsically disordered proteins that can self-assemble into liquid-like droplets. We’re trying to understand the code of protein sequences that can self-assemble in this way, and why, and what are the unique properties of those droplets. These protein droplets form by phase separation, similar to how oil will form droplets in water when a person is cooking in the kitchen. This all happens at the cellular scale, so as you can imagine, these are very small droplets.
What did your experiments show?
We found that the protein code is even more interesting and complex than we previously thought. Everything in the protein sequence matters.
With our collaborators at Texas A&M and the University of Delaware, we tested proteins in which we changed one or a few letters at a time. We found that many different protein sequences can self-assemble into droplets, even several proteins we did not expect to do so. And we found that altering the letters in a protein changes how readily it self-assembles and controls the viscosity of the droplets that the proteins assemble into.
Some letters are more important than others, and interact more strongly with other letters, but all the letters have some contribution to the protein language.
Where are you heading with this work?
This research is a step toward broadening the understanding of cellular biology – to help us better understand how molecules can organize and give rise to functions inside a cell much smaller than the period at the end of this sentence.
It moves us closer to developing treatments for diseases involving pathologic protein aggregates – clumps of proteins that cells can’t break down – like in Alzheimer’s and Parkinson’s. And this knowledge will also help us engineer novel bioengineered materials and biopharmaceuticals.
Other contributors to this research included: Shiv Rekhi, Azamat Rizuan and Jeetain Mittal from Texas A&M University, and Cristobal Garcia Garcia and Kristi Kiick from the University of Delaware.